ACCESS-NRI Intake Catalogue
The ACCESS-NRI Intake Catalogue provides a Python interface to the collection of climate data products that form the ACCESS Data Catalogue on Gadi. For detailed information, Python tutorials and usage examples, please refer to the ACCESS-NRI Intake Catalogue documentation.
Each catalogue entry corresponds to a different data product, where the columns describe the attributes associated with that product (e.g., available models, frequencies and data variables).
Users can filter or search these attributes to find the products relevant to their needs.
For example, a user could search for data products containing variables X, Y and Z at a monthly frequency.
The ACCESS-NRI Intake Catalogue also enables users to directly load the desired data in Python, without having to know the location or structure of the underlying files.
Prerequisites
- Join the xp65 NCI project
The catalogue table files are in theconda/analysis3environment in xp65. Join the xp65 project by requesting membership on its NCI project page. - Join any projects that has the data you're interested in.
The easiest way to use the catalogue is through JupyterLab on the Australian Research Environment(ARE).
Example: using the ACCESS-NRI Intake Catalogue to find, load and plot data with Python
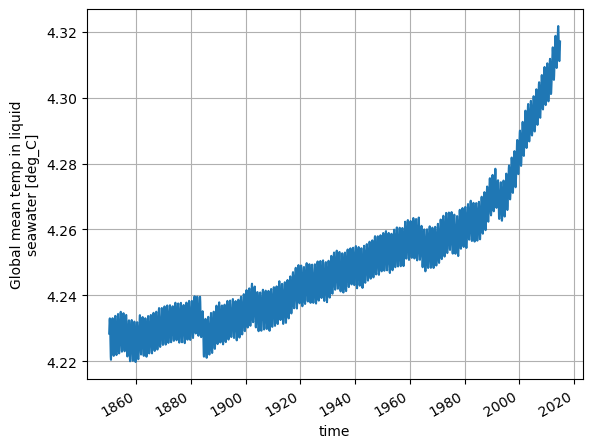
We can plot an ocean variable called temp_global_ave for an ACCESS-ESM1.5 run called HI_CN_05 as follows:
import intake
import matplotlib.pyplot as plt
# Load the catalogue
catalog = intake.cat.access_nri
# This returns an Xarray Dataset
dataset = catalog["HI_CN_05"].search(variable="temp_global_ave").to_dask()
# Plot the data
dataset["temp_global_ave"].plot()
plt.title("")
plt.grid()